About Us
Our research is mainly on the development of eukaryotic genome and RNA editing tools, with the focus on high-throughput functional genomics. The combination of forward and reverse genetic means is employed, often in a high-throughput fashion, for the identification of host genes important for host response during microbial infections or tumorigenesis.
致力于发展基因编辑技术、功能基因组学以及基因治疗;在此基础上研究癌症、感染等重大疾病发生机制,为发展高效治疗手段提供新的药物靶点和思路。



News
MORECell | 魏文胜团队与合作者联合开发新型通用型CAR-T...2025-08-22
2025年8月21日,北京大学魏文胜团队、解放军总医院韩为东团队联合博雅辑因,在Cell发表题为“Glycan shie...
Nature Communications | 成环有道:魏...2025-08-12
2025年8月10日,北京大学/昌平实验室魏文胜团队在Nature Communications发表题为 “Self-s...
祝Wei Lab 2025届各位毕业生们毕业快乐2025-07-03
祝贺实验室14位优秀学子圆满完成学业,其中徐艺源、郑文娟、陈盎、唐慧贤、牛煦然、张巍、沈勇7位同学获得博士学位;莫滨瑞、...
Areas of Focus
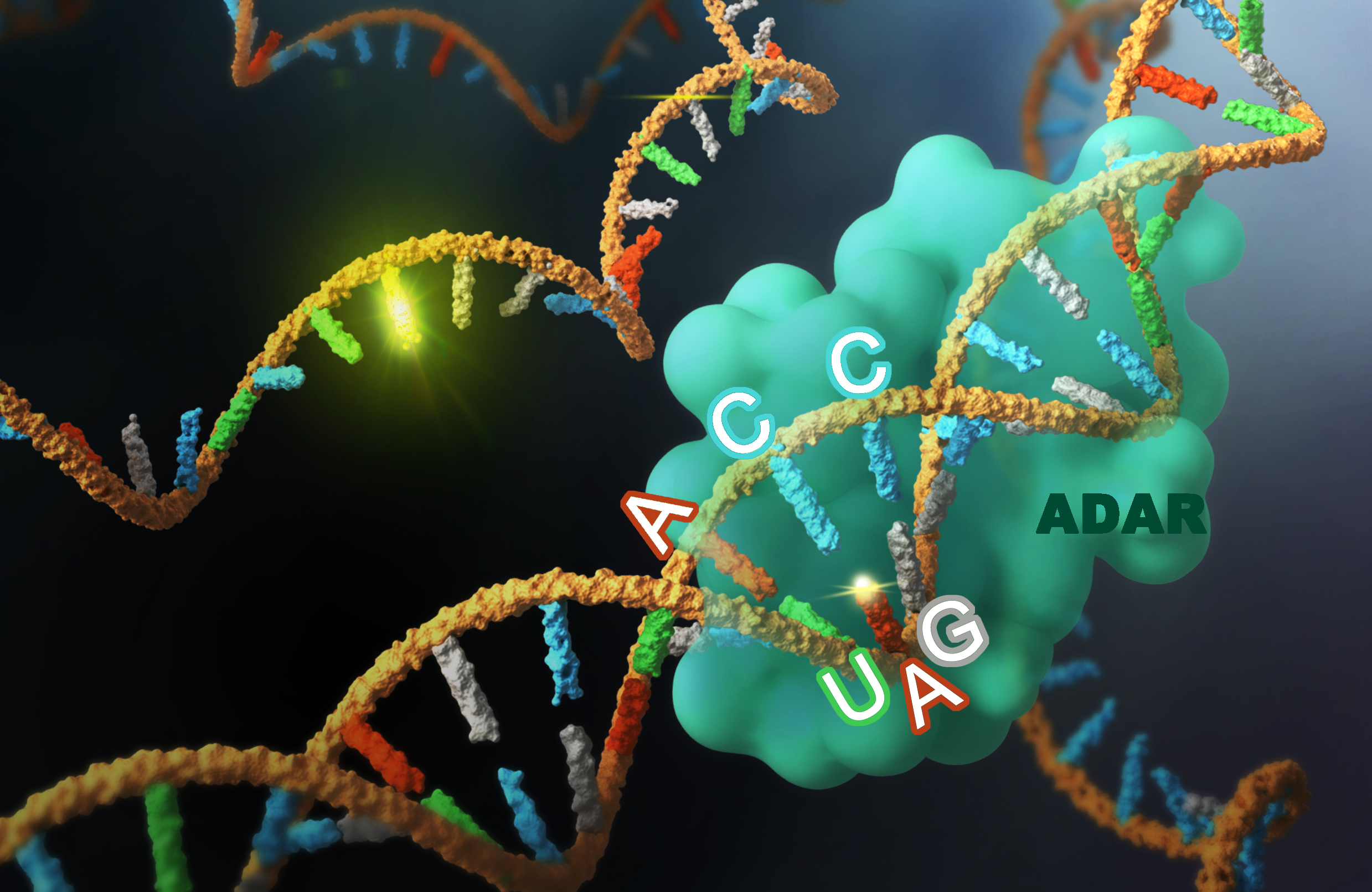
Genome and RNA editing
We are interested in developing novel genome and RNA editing tools and optimizing current editing...
Genome and RNA editing
We are interested in developing novel genome and RNA editing tools and optimizing current editing...

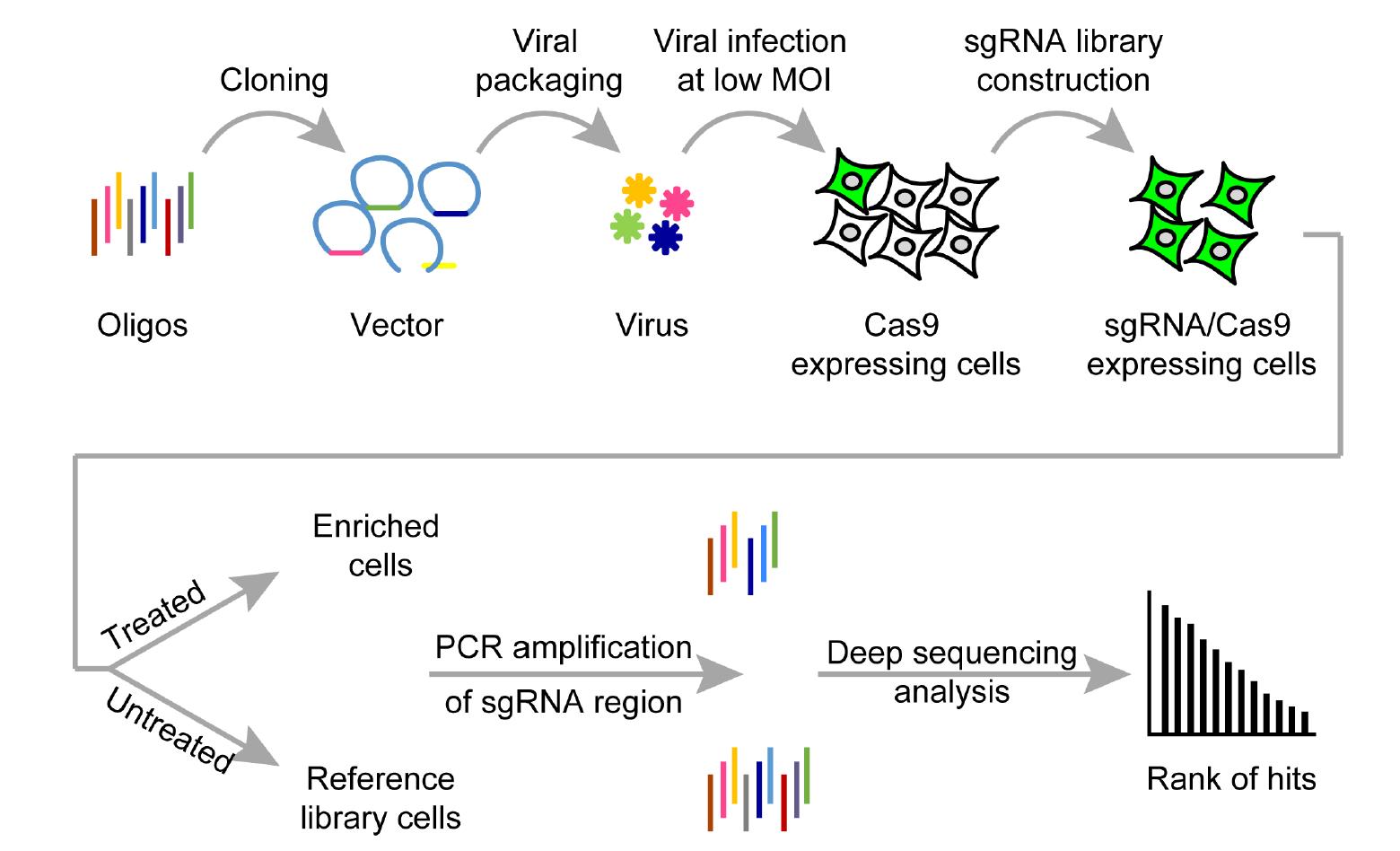
High-throughput functional genomics
We are developing variety of high-throughput screening platforms based on genome and RNA editing ...
People
Publications
- Glycan shielding enables TCR-sufficient allogeneic CAR-T therapy.
- Self-splicing RNA circularization facilitated by intact group I and II introns.
- Huaier suppresses lung cancer by simultaneously and independently inhibiting the antioxidant pathway SLC7A11/GPX4 while enhancing ferritinophagy.
- Prime editor-based high-throughput screening reveals functional synonymous mutations in human cells.
- Super-resolution imaging informed scRNA sequencing analysis reveals the critical role of GDF15 in rejuvenating aged hematopoietic stem cells.
- UNC0638 inhibits SARS-CoV-2 entry by blocking cathepsin L maturation.
- A genome-wide base-editing screen uncovers a pivotal role of paxillin δ ubiquitination in influenza virus infection.
- A mitochondrial disease model is generated and corrected using engineered base editors in rat zygotes.
- Efficient mitochondrial A-to-G base editors for the generation of mitochondrial disease models.
- Massively parallel interrogation of human functional variants modulating cancer immunosurveillance.
- Multi-dimensional bio mass cytometry: simultaneous analysis of cytoplasmic proteins and metabolites on single cells.
- Precise modelling of mitochondrial diseases using optimized mitoBEs.
- N-terminal acetylation of transcription factor LIP induces immune therapy resistance via suppression of PD-L1 expression in non-small cell lung cancer.
- Mapping functional elements of the DNA damage response through base editor screens.
- Protocol for high-throughput screening of functional lysine residues in cell fitness.
- Functional profiling of serine, threonine and tyrosine sites.
- Programmable DNA pyrimidine base editing via engineered uracil-DNA glycosylase.
- Streamlined process for effective and strand-selective mitochondrial base editing using mitoBEs.
- Unsynchronized butyrophilin molecules dictate cancer cell evasion of Vγ9Vδ2 T-cell killing.
- CEA cell adhesion molecule 5 enriches functional human hematopoietic stem cells capable of long-term multi-lineage engraftment.
- Unbiased interrogation of functional lysine residues in human proteome.
- Utilizing AAV-mediated LEAPER 2.0 for programmable RNA editing in non-human primates and nonsense mutation correction in humanized Hurler syndrome mice.
- Strand-selective base editing of human mitochondrial DNA using mitoBEs.
- CRISPR-free, strand-selective mitochondrial DNA base editing using a nickase.
- DddA homolog search and engineering expand sequence compatibility of mitochondrial base editing.
- Human FcRn Is a Two-in-One Attachment-Uncoating Receptor for Echovirus 18.
- Regulatory elements can be essential for maintaining broad chromatin organization and cell viability.
- Circular RNA Vaccines against SARS-CoV-2 and Emerging Variants.
- Gene editing: from technologies to applications in research and beyond.
- Gene editing and its applications in biomedicine.
- Engineered circular ADAR-recruiting RNAs increase the efficiency and fidelity of RNA editing in vitro and in vivo.
- Noncoding loci without epigenomic signals can be essential for maintaining global chromatin organization and cell viability.
- Sensing of cytoplasmic chromatin by cGAS activates innate immune response in SARS-CoV-2 infection.
- Genome-wide CRISPR activation screen identifies candidate receptors for SARS-CoV-2 entry.
- Low-density lipoprotein receptor-related protein 1 is a CROPs-associated receptor for Clostridioides difficile toxin B.
- Genome-wide interrogation of gene functions through base editor screens empowered by barcoded sgRNAs.
- TRIM26 is a critical host factor for HCV replication and contributes to host tropism.
- Activation and evasion of type I interferon responses by SARS-CoV-2.
- Heightened Innate Immune Responses in the Respiratory Tract of COVID-19 Patients.
- Reply to: Fitness effects of CRISPR/Cas9-targeting of long noncoding RNA genes.
- Structural Insights into the Specific Recognition of 5-methylcytosine and 5-hydroxymethylcytosine by TAL Effectors.
- PASTMUS: mapping functional elements at single amino acid resolution in human cells.
- Programmable RNA editing by recruiting endogenous ADAR using engineered RNAs.
- Human Neonatal Fc Receptor Is the Cellular Uncoating Receptor for Enterovirus B.
- In vivo ways to unveil off-targets.
- Adopt a moratorium on heritable genome editing.
- Guide RNAs with embedded barcodes boost CRISPR-pooled screens.
- CRISPR twins: a condemnation from Chinese academic societies.
- Interrogating the noncoding genome in a high-throughput fashion.
- Gene editing technology and its research progress in China.
- Genome-wide screening for functional long noncoding RNAs in human cells by Cas9 targeting of splice sites.
- PrePAIRing Cas9s for screening success.
- A surrogate reporter system for multiplexable evaluation of CRISPR/Cas9 in targeted mutagenesis.
- Attachment and Postattachment Receptors Important for Hepatitis C Virus Infection and Cell-to-Cell Transmission.
- Glucosyltransferase Activity of Clostridium difcile Toxin B Triggers Autophagy-mediated Cell Growth Arrest.
- Live visualization of genomic loci with BiFC-TALE.
- Deciphering TAL effectors for 5-methylcytosine and 5-hydroxymethylcytosine recognition.
- Painting a specific chromosome with CRISPR/Cas9 for live-cell imaging.
- Genome-Wide CRISPR/Cas9 Screening for High-Throughput Functional Genomics in Human Cells.
- Questions about NgAgo.
- Long-term dual-color tracking of genomic loci by modified sgRNAs of the CRISPR/Cas9 system.
- Assembly of Customized TAL Effectors Through Advanced ULtiMATE System.
- Mapping regulatory elements.
- Simultaneous generation of multi-gene knockouts in human cells.
- Making better CRISPR libraries.
- Genome-scale deletion screening of human long non-coding RNAs using a paired-guide RNA CRISPR-Cas9 library.
- Divergent roles of BECN1 in LC3 lipidation and autophagosomal function.
- High-throughput screens in mammalian cells using the CRISPR-Cas9 system.
- A Dual-Reporter System for Real-Time Monitoring and High-throughput CRISPR/Cas9 Library Screening of the Hepatitis C Virus.
- A microfluidic live cell assay to study anthrax toxin induced cell lethality assisted by conditioned medium.
- Chondroitin sulfate proteoglycan 4 functions as the cellular receptor for Clostridium difficile toxin B.
- Bidirectional effect of Wnt signaling antagonist DKK1 on the modulation of anthrax toxin uptake.
- Complete decoding of TAL effectors for DNA recognition.
- High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells.
- ULtiMATE System for Rapid Assembly of CustomizedTAL Effectors.




